Endogamy Part II – One Segment from One Ancestor
Review
In Endogamy Part I (Shared DNA), we found that the total cM shared between you and a Match is multiplied by the number of times you had the Common Ancestor (CA) in your Tree. So if you and your Match were 5th cousins (5C), you would normally share 3.4cM. If your CA (between you and your Match) was in your Tree five times you would tend to share, on average, 5 x 3.4cM = 17cM. If your Match has that CA in her Tree three times, the total you would tend to share, on average, would be 3 x 5 x 3.4cM or 51cM. For close-cousin Matches (say 1C, 2C, 3C), this make a big difference. For distant-cousin Matches (say 6C-8C or more), where you would probably not match at all most of the time, the endogamy may increase the total cM enough that you’ll actually get an above-threshold Match. Note: a 7C would normally share 0.1cM, so even with an Endogamy factor of E15, you’d only share 1.5cM, on average, and you’d need to be on the tail of the distribution curve to exceed a 7cM threshold for a Match.
In this post – Endogamy PART II – we’ll look at what happens to an individual segment.
Ground Rules
Shared Segment means an IBD segment, from an Ancestor. I usually consider all shared segments over 7cM in a Triangulated Group to be IBD segments.
One Ancestor or one CA means one Ancestral line. See CA and MRCA for a discussion of the CA for a shared segment. Note: at different cousinship levels, there may be intermediate CAs, or MRCAs, which are all in one Ancestral line. The shared segment comes from the most distant CA of that Ancestral line. In other words, all the Matches with shared segments (think a Triangulated Group), will have a CA with you on one Ancestral line – you and they will all descend from one CA.
Endogamy means a CA is in our Tree multiple times. So Ancestor A can be represented by A1, A2, A3, etc. for each time that Ancestor is in out Tree. As you read on, you’ll note that it’s important to treat each one (A1, A2, A3, etc.) as a separate Ancestor (even though they are all the same individual).
Assumption: Each Match will have one CA with you. I know in many cases a Match may share multiple Ancestors with you. But for the purposes of this blog post, we will only look at the effects of one CA. We have to build up, one concept at a time. Learn the concept in this post.
Shared Segments from Duplicate Ancestors – analysis
Let’s look at a shared segment between you and a Match. It could be any shared segment. Just to add some reality to this discussion, let’s say it’s on Chr 10 from 8 to 20Mbp with 23cM. We’ll call this SEG-1.
In this example, the CA for you and a Match is in your Tree twice: A1 and A2. Both A1 and A2 are the same person, so both A1 and A2 have the same SEG-1. Let’s look at Figure 1 and see how far SEG-1 can descend toward you.
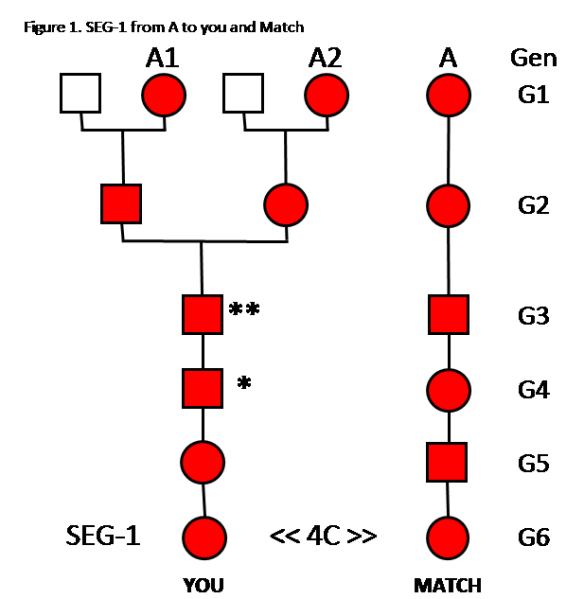 Analysis of Figure 1:
Analysis of Figure 1:
In Generation 6 (G6), YOU and a MATCH are 4C, and share SEG-1. SEG-1 came from Common Ancestor A. Red indicates that person has SEG-1.
In G1, your 2 ancestors, A1 and A2, have SEG-1; and your Match’s ancestor, A, has SEG-1. These are all the same individual – so naturally she has SEG-1, no matter where she is in your Tree or your Match’s Tree.
SEG-1 is passed down from A to the Match through one ancestral line.
In G2, SEG-1 is passed from A1 to her son; and from A2 to her daughter.
In G3 A1’s son passes SEG-1 to the paternal chromosome in his son; and A2’s daughter passes SEG-1 to the maternal chromosome in her son. This G3 son now has two copies of SEG-1, one on his paternal Chr 10, and one on his maternal Chr 10. This is indicated by **.
In G3, the ** father, recombines his two Chr 10s to make one Chr 10 to pass to his son. Only one of the two SEG-1s can be passed on. The G4 son will only get one SEG-1. This is indicated by *. We don’t know whether the maternal or paternal SEG-1 is passed on – it’s a 50/50 chance for either. If you need a refresher on recombination and how only one area of two chromosomes can be passed down, please review Segments: Top-Down.
In G4 and G5, SEG-1 will be passed down to you, and you will share this segment with your 4C Match.
Starting with G1, there are lots of possibilities, but for you and your Match to both share SEG-1, it has to start in a CA (in this case A, A1 and A2) and be passed down through each generation to you and your Match.
One Segment from One Ancestor
This is a fundamental concept of genetic genealogy. Each shared segment can come from only one Ancestor. This means from only one of several Ancestors (A1, A2, A3, etc.) in Trees with endogamy. This can be extended to each Triangulated Group can come from only one Ancestor. A corollary is that a different shared segment (or TG) can come from a different Ancestor. With an Ancestor in your Tree 5 times, it is possible for you to have a different segment from each one. It’s also possible for one Ancestor to pass down several different shared segments (in different TGs). So although we can say “One Segment is from One Ancestor”, the reverse is not true. We have to say “One Ancestor can pass down Multiple Segments” (or no segment).
Shared Segments from Multiple Ancestors – analysis
We can apply this same concept – One Segment from One Ancestor – to even greater endogamy. See Figure 2.
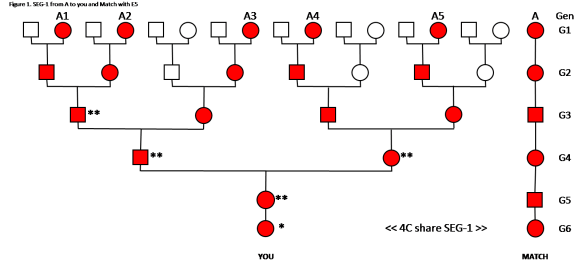
Analysis of Figure 2:
In this case we have E5, the Common Ancestor is in your Tree five times. When cousins (descending from the CA) marry, their child may get two SEG-1s – one on each chromosome. This is indicated by the double **. The next generation gets only one SEG-1 from that line. As noted with the A3 line, the son in G4, could have double ** – one from A1 or A2 (paternal chromosome) and one from A3 (maternal chromosome). The daughter is G5 got a paternal SEG-1 from A1, A2 or A3, and a maternal SEG-1 from A4 or A5. You got a SEG-1 from A1 or A2 or A3 or A4 or A5. At this point we cannot tell which of your Ancestors passed down the SEG-1, but we do know it could only have come from one of them.
In this discussion, I’ve used the “worst case” scenario – each of your A Ancestors passed SEG-1 down as far as she could. In fact, SEG-1 is subject to the 50/50 rule – half the time it will be passed down, half the time it won’t. However, the fact that we share SEG-1 with a Match, and have a Common Ancestor A with that Match, means at least one of your Ancestors A had to pass it all the way down to you and the Match.
We are all well aware that you and a Match probably have multiple Common Ancestors. Again, this analysis does not sort out which Ancestor is the correct Common Ancestor for SEG-1, nor does it sort out which one of the multiple Common Ancestors it is. This analysis just establishes the point that:
One Segment comes from One Ancestor
A very unusual exception
In the case where your parents are cousins, it is possible (but not very probable), that you would carry two SEG-1s (from this example) – one paternal, one maternal. This would be the case in Figure 2 if you were the daughter ** in G5. At GEDmatch, any such segment areas would be highlighted with their “Are Your Parents Related?” tool. So it’s easy to check for such segments, and be aware of where they exist (Chromosome and Start/End locations). In all other cases you don’t need to worry about this issue. For any shared segments meeting this very unusual criteria (exactly the same segment on both chromosomes), you wouldn’t know which of your two ancestors it came from. If there were any difference in these two shared segments (they were not exactly the same), then chromosome mapping would usually tell you which one was which.
Summay:
Each shared segment comes from one Ancestor. In the case of endogamy with multiple identical Ancestors, each shared segment comes from only one of them.
It’s possible to have ten identical ancestors in your Tree, and to get a different shared segment (as in a different TG) from each of them.
16E Segment-ology: Endogamy PART II – One Segment from One Ancestor by Jim Bartlett 20160104

Hi Jim
One scenario I don’t see in your examples is my case from the Isle of Lewis in Scotland (like many isolated communities where cousin marriages are relatively common) – my grandfather (age 50) married his 1C1R (age 23). So there’s a generation ‘dilution’ of the doubled-up DNA, which I calculate makes a E1.5 factor. This is approximately borne out by readings from 2 first cousins (factor of 1.3 and 1.2) and 3 second cousins (2.2, 1.6 and 1.3). I’m getting three more cousins tested so that we can refine the figures. The reason is we have a closely related mystery cousin and we’re trying to work out how much to discount the readings due to endogamy.
This is not an easy topic!
Cheers, Mike
LikeLike
Mike, three things:
1. I’m happy to see the E factor being used and validated;>)
2. Only use an E factor when the Common Ancestor is there. If some other CA turns out to be the correct one, then calculate the E on that Ancestor. In other words an E factor does not include endogamy in multiple lines.
3. As even in no endogamy cases, the shared cM can usually come a variety of cousinships; the same is true with an E factor: there are usually several possibilities – the smaller the cM, the more possibilities.
LikeLike
My dad is Cajun and his family tree looks very similar to your E5 example. The main difference is that his paternal line is one generation closer than the other lines. So Joseph Landry is his 2nd great grandfather as well as his 3rd great grandfather four times. And not only that, Joseph was related to his wife 8 different ways (3C, 3C1R…). So when I tested my dad at 23andMe, he ended up with about a thousand ‘2nd and 3rd’ cousins according to the amount of matching DNA. When I was able to find the MRCA, the match usually ended up being a 7C or 7C1R. By the time I get that far back, there are several other MRCA that I can find. (And many undetermined which are probably CA as well.) And I have at least one ancestor that shows up in my tree 30 times.
It is complicated enough when you are looking at just one CA and finding the ways they are connected in your E5 example. For my father, by the time you go back as far as you show in your example, everyone in his father’s tree is included in his mother’s family tree. So you may be able to determine who the segment came from (how, I don’t know), but you wouldn’t know what route it took.
Sadly, I do not spend any time trying to untangle the DNA results of my dad and his matches. Fortunately for me he spent 40 or so years untangling the paper trail and tracing most of his lines back to Acadie.
LikeLike
Van, The blog post was just an illustration of endogamy. I agree that it can get very complicated. The point is that with endogamy, we cannot trust the total amount of DNA as an indicator of how close a Match/cousin is – we are far better off focusing on the largest individual shared segment.
The only way I know of to tell which line the DNA came from is through luck – finding a cousin who relates to one line, but not the others – not always possible…
Jim
LikeLike
Seth….that’s a decent amount! You would share common ancestors further back probably beyond & further back, possibly multiple times and/or you have more than one common ancestor.
LikeLike
Seth – you and your 3rd cousin may well share another Common Ancestor, and/or you may have gotten some “endogamy” cMs from either of these.
LikeLike
There is something very basic that I’m failing to understand about figure 1. Since A1 nnd A2 are the same person it looks like siblings are marrying in G2. I don’t see any other way that G3 could get the same segment from both his maternal and paternal lines but I’m sure that a sibling marriage is not what is being suggested here. What am I missing?
LikeLike
June, you are correct. I goofed on the figure and should have added another generation near the top. I’ll fix it so that it’s first cousins marrying each other. The point of the blog post is that the shared segment at the bottom of the figure can only come down one path – from one of the identical ancestors. Sorry for my error in the figure.
Jim – http://www.segmentology.org
>
LikeLike
But in truly endogamous populations, wouldn’t both people have that same ancestor multiple times in a tree? Or even multiple common ancestors multiple times for both?
LikeLike
In my experience, yes, definitely. I just mentioned that I know of a couple of people descending from just 12 people (one of them from all 12) that founded a population and indicated the number of times one of them descend from all 12.
LikeLike
Yes to Q1. In any population both you and your Match will have multiple ancestors in your Trees. And in endogamous populations, this is even more pronounced in a genealogical timeframe. I noted, and quantified, this in Engogamy PART I.
And yes to Q2. The point of this blogpost is two fold: one is you and your Match get the (usually one) shared segment from a Common Ancestor; and that shared segment came from only one instance of that Common Ancestor.
I’m not trying to trivialize the problems with endogamy, but I am trying to establish some fundamental concepts that can help us sort out endogamy. IMO, there is too much loose talk about “population” segments, and how intertwined endogamous populations are. I want to tease apart some of these statements – to focus on some concepts that we can work with – break it down so we can analyze each specific case. Endogamy PARTS I and II, establish specific concepts we can use, ways we can better communicate about endogamy. At least that is my goal.
LikeLike
I’m trying to see if I can predict correctly the amount of total shared two people can have coming from a small founding population of 12 people in 1790. 6 men and 6 women. But so far, if I even try to come closely to what you are saying, it doesn’t seem to add up, probably because for the women alone they were from an endogamous group.
The person I am comparing descends from two of the women 10 times, and one of the men 8 times, and all the others between 2 – 6 times each. Out of those women, only one had 1 husband, 3 of them shared 2 husbands and two of them had 3 husbands. 3 of the men had at least 2 wives, while some of the others had more or less.
I really didn’t focus too much on the matching segments only because they are from a small founding population, it was easy to keep track of their genealogy. But these people match a bunch of us who have no ties to the island that they are from.
You can read more about it here:
LikeLike
Read Endogamy PART I. Endogamy only increases the cM when the Common Ancestor. There isn’t a way (that I know of) to calculate a general increase in cM based on all the endogamy. It’s specific to the Common Ancestor.
Jim – http://www.segmentology.org
>
LikeLike
I commented on Endogamy PART I back on Dec 2nd.
LikeLike
Trying to predict such numbers with a usable accuracy may be futile as the variance of the predictions, even if mathematically sound, will be so high as to make them useless.
As an example of one of many difficulties, in human male meosis the literature indicates that there is an average of 1.1 to 1.2 crossover events per autosomal chromosome while it is 1.7 to 2.2 in creation of female ova. The implication is that segments inherited from an individual down a strictly male line will differ in size from those inherited down a strictly female or mixed sex line.
There might be some hope of getting some benefit from a simulation.
LikeLike
Grumpy, here’s the deal. The literature actually shows there are an average of 27 crossovers for a male and 41 for a female. And they are spread over 22 or 23 chromosomes, respectively (not on Chr Y). You cannot divide by the number of chromosomes to get an average per chromosome – instead, you may divide by the number of Mbp or cM in an entire genome to get crossovers per Mbp or per cM. This means longer chromosomes get more crossovers than shorter ones do.
And each recombination event is recombining a paternal and a maternal chromosome, so there is really not much difference between all-male and all-female lines.
LikeLike