This question comes up often. The answer is: we cannot tell from just the fact that two shared DNA segments overlap in a chromosome browser. Here is the picture we see:

In this picture, you are normally A and you have two Matches, B and C, which show as overlapping on Chromosome 6. Because they overlap, is this Triangulation? Do A, B and C shared the same Common Ancestor? We cannot tell from this picture.
Assuming the shared DNA segments are Identical By Descent (IBD) – generally true for all such shared segments over 15cM – there are two possibilities:
- They are on different Chromosome 06’s in A. Remember we have two of each Chromosome – one from our mother and one from our father.
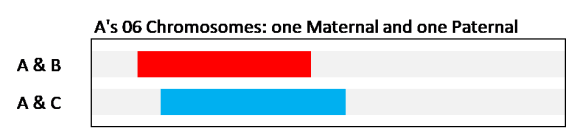
In this case, we are (somehow) looking at just A’s two Chromosome 06’s and showing where the shared DNA segments are on A’s DNA. It looks just like the picture we saw in the browser – two overlapping DNA segments. But in this case A & B are sharing on A’s maternal Chromosome 06; and A & C are sharing on A’s paternal Chromosome 06. These two Chromosome 06’s are physically separate (think of two strands of spaghetti). Because A & B have a shared DNA segment, they have a Common Ancestor (CA) who passed that DNA down to them. Because we know in this example that it’s on the maternal Chromosome (the one from A’s mother), we know the CA is on A’s maternal side. Similarly, we know the CA with C is on A’s paternal side. Yes, there is a very unlikely chance that these two CA’s could be the same person, and the DNA segment came down two very different paths to A’s mother and father. I’ll not be sarcastic here – you can decide for yourself if you think that is possible (or what the probability is) in your case.* In general, in genetic genealogy, we conclude that B & C are probably not related to each other – at least not on this segment.
- Alternatively, the two shared segments are on the same Chromosome 06 in A – let’s say, for example, they are both on the maternal side (imagine the two bars below on one Chromosome).
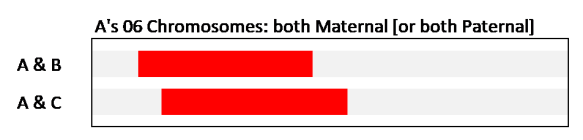

In this case, we are (somehow) looking at just A’s one maternal Chromosome 06, and showing where the shared DNA segments are. Again, it looks just like the picture we saw in the browser – two overlapping DNA segments. But in this case A & B and A & C are sharing on A’s maternal Chromosome 06 (they are both on the same strand of spaghetti). From the beginning of the A & C shared segment to the end of the A & B shared segment, we are looking at the exact same place on A’s Chromosome 06. For there to be a match, all the tested markers (SNPs) are the same. In general, in genetic genealogy, we take this to mean that this DNA came from the same Common Ancestor. It came from that CA down to A and to B and to C. Because both B and C share this same segment of DNA found on one Chromosome 06 in A, both and B and C should themselves show up as a Match to each other. After all they have the same DNA over this area of their own Chromosome 06.
You may have noticed that I stated each explanation of the two possibilities with: “In this case, we are (somehow) looking at…” Well we can’t just look at just one chromosome in a browser and compare it to someone else’s DNA. We don’t have that technology for genealogy DNA testing. But if we could, that is what we would see (probably without the color coding). But we cannot! We can only visualize it. So what can we do?
We use reverse logic. In the first possibility, we noted that B & C wouldn’t match each other; and in the second possibility, we noted that B & C should match each other. That is information we often can determine (at 23andMe, MyHeritage and GEDmatch – and round-aboutly at FTDNA). So, we say that if A matches B, and A matches C on the same/overlapping DNA segment, AND B matches C there too, it indicates the second possibility above – the three of them share the same Common Ancestor. This case of A=B=C=A is called segment Triangulation, and the three Matches are in a Triangulated Group [TG]. There is more about Triangulation here.
In my case, I have close to 20,000 Match/segments – each shared DNA segment is in one of 372 TGs which cover all of my DNA. In other words, these 372 TGs form a segment map of my 45 Chromosomes. The objective now is to determine the Ancestors who passed these TGs down to my parents and then to me.
*If you want to check to see if you have the same segment from your mother and your father, upload your DNA to http://www.GEDmatch.com and use the “Are Your Parents Related” program. It will show you any such segments, which is good information to have in any case.
[11D] Segment-ology: Are Overlapping Segments Triangulated? by Jim Bartlett 20200414

Hi Jim,
I’m doing a catch-up on your blog, so please forgive if the answer is elsewhere. Just point the way…
Whether B matches C can be determined “… round-aboutly at FTDNA” — can you give me a hint here? I am reasonably adept at triangulation at the other companies you mentioned but have yet to figure out FTDNA.
Thanks for the sharing your wealth.
Kate
LikeLike
Kate – at FTDNA, if two Matches overlap you and each other AND each one is in the other’s ICW list, that is sufficient to Triangulate those segments. It’s basically 99.99% if that happens with 3 Matches. If ever there is some doubt a shared Segment being in a TG, check to see there are ICW Matches that are in the same segment on the other side. In other words if you have A, B, C, and D in a Paternal TG at the beginning of Chr 04; and C looks suspicious (for any reason); then look at the ICW list with C and see if any of those Matches are in your Maternal TG at the beginning of Chr 04. The Matches with shared DNA segments at the beginning of Chr 04 (or any other segment), should all match each other on one side or the other – they will form TGs on both sides and that would be reflected in their ICW lists. Jim
LikeLike
Thank you very much, Jim. I have understood the concept. It takes a bit more work with endogamy, I believe, because the ICW lists are largely common. Still, I’m working the strategy of seeking out the non-ICWs and making TGs. Great tip! Thank you very much!
LikeLike
Jim, thank you for linking back to this page. It illustrates my scenario exactly. The 2nd figure (where both bars are red) shows A&B, and A&C as a triangulated group. A matches B, A matches C, and B matches C. The post summarizes: “The three of them share the same Common Ancestor (CA). This case of A=B=C=A is called segment Triangulation, and the three Matches are in a Triangulated Group [TG].” Assuming identical by descent (IBD), doesn’t this imply that all the SNPs from the left edge of A&B to the the right edge of A&C descend from that ancestor?
LikeLike
Andy, Yes – assuming IBD. However, this situation begs for more data. Somewhere, there should be a shared DNA segment that will span the whole area – assuming it’s from one Common Ancestor. Of concern is the fairly small overlap for your B to C Triangulation. One the other hand, it’s OK to assume you have one TG. What’s the harm? You still need to find the Common Ancestor; and you really should have other corroborating Matches. My real concern is confirmation bias. If this is just a Triangulation exercise, that’s fine – every segment needs to go somewhere: a paternal TG, a maternal TG, or the IBC junkpile. However, if someone starts with a particular solution in mind, and is searching for scraps of evidence to “fit” (what I call: trying to put on Cinderella’s slipper), it concerns me. If we get one segment in a TG wrong – no big deal – we’ll search and probably never find a Common Ancestor with that Match. There is always a chance, with under 15cM shared DNA segments, that some Match will have an IBC “fit” to the TG. Not a big deal, unless that Match is the lynchpin to a “proof”. That’s why we should always have a preponderance of evidence for a “proof”. Collecting and Triangulating segments, is a way to group Matches. The CA for each TG needs to have multiple pieces of agreement. Jim
LikeLike
Got it. Thank you. If one TG with just two matches were all that’s available to confirm an ancestor, it would be shaky ground indeed. The small overlap of the two matches was contrived; I was just trying to clarify the definition of TG, which you have resolved for me. Ideally, there will be multiple matches within a TG, and they will span the entire ancestral segment. I’m finding this is the case with many of the TG’s on my paternal Georgia side. On the maternal side (Germany), the data are sparse, like the hypothetical example. There are practically no TG’s of any length & depth at the online database I’ve been working. Thank you again for the added clarification.
LikeLike
Andy, Thanks for the feedback. My maternal grandmother, had Scot father and German mother, whose parents were immigrants n 1860 timeframe – I have a handful of 2C and 3C from US, and almost nothing 4C and beyond (ancestors in the old countries). I, too, have a few small gaps in the TG map on this part of my ancestry.
LikeLike
Jim, Inspired by your example, I want to learn how to use Excel to manage my Ancestry A-C results. Have never use spread sheets before so would appreciate advice in getting started:
1)which version of Excel would you suggest to do what is needed (i.e., to do it the way you are doing it). I have a Chrome Book, also a HP laptop w/Intel Pentium Processor, if that is needful info here).
2) can you recommend a site for learning Excel basics? I am 77 & don’t consider myself a technophile AT ALL, altho that hasn’t kept me from using computers to organize my Genealogy DB (now with over 1/2 million names in Brother’s Keeper) for several decades now. Any insights or advice you can share would be greatly appreciated!
LikeLike
Carolyn,
I subscribe to Microsoft 365, which includes Excel and Word and Power Point – all of which I’ve used for more than 20 years. If you are serious about starting to learn Excel to track DNA segments, I would recommend a local Community college class, or tutor – maybe using Zoom. What I do with spreadsheets is probably in the intermediate range….
Another thought is to study your Shared Matches, and create Notes about what you find. I have gone through all of my ThruLines Matches and type the Common Ancestor(s) into the Note boxes for each one. After I’ve done that for all ThruLines Matches (about 1,000 of them in my case), I can then look at their Shared Matches and note if there is a clear trend – most of the Shared Matches being on the same line. When this happens, I focus on that line (and the collateral lines) and look at the other Shared Matches and their Trees for hints that would tie them to that line. It’s almost like magic sometimes..
The Clustering programs present you with a spreadsheet result – so all you have to do is scroll down the spreadsheet and look at the Cluster groups – which groups are unually on the same Ancestral line. I did a blog post on the three popular Clustering programs.
Hope this helps, Jim
LikeLike
Douglas – I’ll offer several thoughts.
1. Everyone’s Ancestry, geography and Matches will be different. I cannot change that – you have to deal with what you get. However, one important step anyone can take is to test at all 3 of the companies which show shared segments (23andMe, FTDNA and MyHeritage), and upload to GEDmatch (where you can ask for up to 100,000 Matches).
2. I’ve formed Triangulated Groups [TGs] that cover all of my DNA – all 45 chromosomes (not the Y). About 10 of them are small gaps between other TGs with no shared segments in them. That means the average size of each TG is about 20cM – some smaller, some larger. Most of the Matches in each TG will overlap most of the rest – to varying degrees.
3. No one really knows (or has published in a peer review journal) how small a shared segment can be used for Triangulation to work 100% of the time or even say 90% of the time. Most of the companies use a lower threshold of 6 or 7cM for their matching, so I have use 7cM as my lower limit for shared segments and for the overlap in Triangulation.
4. TGs should initially be formed from the large segments – preferrably with 15cM overlaps (15cM is generally accepted as the a threshold for virtually 100% IBD segments). After all the large segments are used, drop down a few cMs and add in more segments, etc.
5. See my blogpost for Anatomy of a TG – not all shared segments in a TG have to overlap all the others – but each shared segment has to match some of the shared segments in a TG.
6. In segment Triangulation we are looking for about 300 to 350 crossover points (my estimate), which paint the picture of how we inherit different segments from different Ancestors.
7. Every IBD shared segment has to fit in somewhere – on one side (chromosome) or the other. If a shared DNA segment clearly does NOT fit on one side, it must be on the other side if it’s IBD.
8. In the early days of atDNA, it took me about two years to map most of my DNA – it wasn’t easy, and there were gaps until more Match/segments came in. Maybe in some cases (like yours?), there just isn’t enough information to do Triangulation on a full scale. It’s not your fault, and it’s not the fault of the Triangulation process.
9. Speaking personally from my experience, I would not adopt a 2cM or 4cM or 6cM overlap rule – there is too much at risk, IMO. Instead, get more data. If you are painting a wall with only a small amount of paint – I wouldn’t try to lightly cover the wall, I’d get more paint and do it right.
10. Each of us has fixed crossover points (which define the start and end of TGs) in our DNA since conception. We are trying to use the data to determine the location of those points. Then the chore is to find Common Ancestors to go with each TG (from the Matches in each TG).
Hope this helps, Jim
LikeLike
As always, thanks Jim for your insights. I simply do not get so many large segment matches and so almost my entire triangularization effort focuses on multiple overlapping IBD matches all in the 3 to 8 cM range… let’s call them clusters. I’ve started using clustering tools instead of TGs to see if more clarity falls out.
Question 1: so if I have a cluster of say 7 testers and I have filtered matching segment sizes to > 10cM and determined that all of the 7 testers match each other over a specific chromosome range, but all of the checkered overlaps are 3 to 8 cM in size, are you saying that this may not be a reliable TG?
Question 2: if you had a smaller number of 1st thru 4th cousins, do you think you could still sort out those hundreds of crossovers from your 18th and 17th century ancestors, based on your considerable experience at this?
BTW, I do have tests at 4 orgs: FTDNA, ANCESTRY, 23andME and MyHeritage – but trying to maintain a master file that keeps all 4 sets of matches organized is very daunting. And BTW it is not just my own DNA but my mother’s, father’s sister, and a few cousins that I have tested. Just way too much data to process, organize and update. I’m impressed with the work you have done, but I simply can’t imagine how you pull it all together into something meaningful and keep it managed with updates. The numbers of matches are many tens of thousands and I find even excel spreadsheets become very difficult to manage when there are so many records. Any hints?
LikeLike
Douglas
Q1 – it probably is OK. The “proof” comes when you form a TG on the other side, adn none of your 7 match anyone in the new TG spanning the same area.
Q2 – It’s hard to say, but I’ll give you the benefit and say it looks like your situation is much harder than mine.
I have about 20 kits that I manage, but I am only working on TGs for one – mine. I tried to do it for my Dad and brother and uncle – but it’s too much. A lot depends on your objective. My objective is to map my DNA to my Ancestors – which Ancestor gave me which genes…. I’m not sure what the objective is for folks who try to keep up with 4 or 40 kits. I did genealogy without atDNA for 35 years. I’m now trying to like my genealogy to specific segment of DNA. My closer relatives help me – to a certian distance – on some TGs, but I’m trying to push the TGs out about 9 generations. It keeps me busy.
Jim
LikeLike
Jim you show an example where the triangulated portion (ie the overlapping portion) is substantial – ie if the segments were 15 cM long, the shared portion is maybe 12 or 13 of the 15cM. But what about the case were the overlap is only 6 or 5 or 4 of 15cM? This is where I get bogged down in triangularization, as strong matching segments only overlap at significantly less percentage – say less than 67% of the largest match. This is because my tree has small-sized families for the past handful of generations, and then before that the colonial families can be large. Hence I have virtually no 2nd and 3rd cousins, not a great deal of 4th cousins, but thousands of 5th thru 8th+ cousins. My segments overlap all over the place, but the overlaps are not 15cM but rather 2 to 6 or 7cM. Its a dog’s breakfast to look at and sort out… especially when a few NPEs are mixed in and so trees don’t align.
LikeLike
Gedmatch tier 1 shows me cross-matched triangulated segments on chromosomes; MyHeritage also shows triangulated matches by segment and chromosome. Are these matches questionable?
LikeLike
Linda – sorry I missed this question months ago. The short answer is that the triangulated segments shown at MyHeritage are correctly Triangulated; Triangulated segments at GEDmatch are correct. And… at 23andMe > Relatives in Common > with a Yes in the SharedDNA column also indicates a correct segment Triangulation (but you need to click on the Yes to see which segments). I do not question any of those. Jim
LikeLike