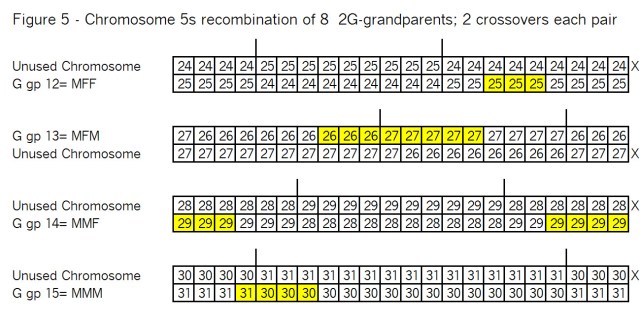
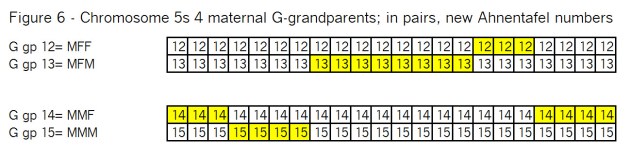
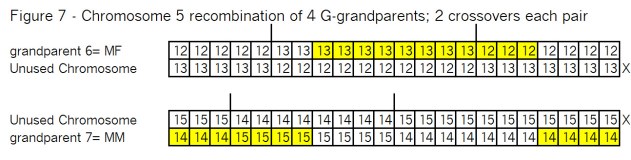
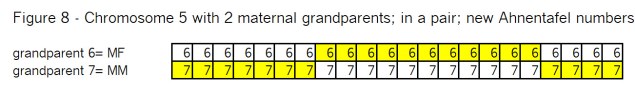
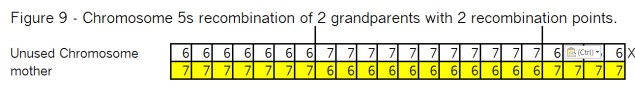
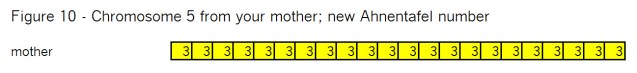
What’s all the buzz about “pile-ups”? In my mind there are three kinds of pile-ups: small, medium and large. They are different, so it’s important to understand each one. In this case Goldilocks should prefer the large pile-ups, but let me go through my views of all three kinds.
Alert: This post contains my opinions about small pile-ups and AncestryDNA (based on my own experience) so you should make your own judgments.
Background
I think the two keys to success with autosomal DNA lie in a robust Tree (as many ancestors out to 13 generations as possible) and as many Match-segments as possible (including as many close relatives as you can get). I spent about a year expanding my Tree as best as I could, and then posted that GEDcom in several places. I’ve tested at all three companies and use GEDmatch. I put every single shared segment I can find over 7cM into my spreadsheet, and I periodically run a Quality Control check against a fresh download to pick up any missed Matches or segments. I currently have 5,000 different individuals with segment data in my spreadsheet, and have determined a Common Ancestor (CA) with 309 of them.
I have compared virtually every segment against other overlapping segments, and formed Triangulated Groups (TGs) that cover over 90% of my 45 chromosomes. It is now rare for me to get a new shared segment that changes my chromosome map in any way. This process has provided some insights on medium and large pile-ups.
Pile-ups
My definition of pile-up sizes:
- Small is smaller than 5cM
- Medium is 5-10cM
- Large is greater than 10cM
Small pile-ups – by my definition, these pileups are composed almost entirely of IBS shared segments. When AncestryDNA first rolled out their autosomal DNA test, their threshold was 5Mbp. This threshold included many shared segments well below 5cM, and resulted in many thousands of bogus Matches. To their credit, they provided a caution about these. When AncestryDNA revised their threshold to 5cM, many of these Matches went away. Part of their explanation was the elimination of “pile-ups”. I agree that these “small pile-ups” should be eliminated. And when they reset their threshold to 5cM, that should have eliminated this problem. However, their explanations continue to stress the elimination of “pile-ups”. I just hope they don’t also toss out Matches in larger pile-ups – throwing the baby out with the bath water.
Medium pile-ups – 5-10cM range. As I gathered as many segments over 5cM as I could and sorted them in my spreadsheet, I noticed a few areas that had many such segments, all in a very narrow chromosome area. Very clearly a pile-up! Virtually none of them matched each other, although they had almost the same segment start/end locations. And there were a lot of them – many more than in large TGs.
In discussions on various email lists, we compared notes, and found that most of these areas were unique to our own experience. In general they were not due to some common feature of most human genomes. A notable exception to this blanket statement is the HLA Region on Chromosome 6 – roughly from 29.8 to 33.1Mbp.
However, most of the other areas were not tied to known issues like the HLA Region. In my analysis, it was not possible for me to link these to one parental side or the other. The fact that these areas include so many IBC segments indicates to me that it’s the combination of both of my chromosomes (maternal and paternal) that allows the “matches”. It’s the unique combination of alleles in these small stretches of DNA that make matching much easier. And this unique combination is only in my genome. On chromosome 18, I have 307 segments in the 7 to 11 cM range. They are all in a very tight area: from location 5,800,000 to 8,700,000bp. Very few of them triangulate.
Sometimes the pile-up area has been documented. On chromosome 15, I have 281 segments in the 7 to 10cM range. They are at: 24,000,000 to 28,000,000 bp. This area partly overlaps a known pile-up area (20,100,000 to 25,200,000). But the known pile-up area is only partly the cause in my case. See 14 small pile-up areas found by Li et al (2014), listed at the ISOGG Wiki: http://www.isogg.org/wiki/Identical_by_descent These medium pile-up areas, and a few others in my experience, are characterized by a very tall pile-up of many segments about the same size in a narrow area just a little larger than the segments. The Li et al (2014) article refers to “regions where excess IBD is detected…” Virtually all of the segments I have noted above are IBS/IBC – they do NOT triangulate with the other segments. A few segments in these regions do triangulate with known close relatives, and each other. I’ve kept those segments in maternal and paternal TGs, as appropriate, covering that area. After all, both my mother and father gave me those areas, and they in turn got them from their parents, etc. It is very probable that these segments are IBD and come from a CA.
My experience is that these are areas with a lot of shared segments in the 7-10cM range that are in a tight area, usually just 10cM wide, and a very high proportion of these segments are IBS/IBC. A few segments in these areas will be IBD, but they will tend to be larger than the 7-10cM segments.
My bottom line for these pile-ups: Unless you have a lot of free time, skip over these areas – particularly the shared segments under 10cM. Concentrate on triangulating any larger segments in these areas and then move on to other areas.
Large pile-ups – these are my favorites. Larger shared segments (over 10cM) that spread out and overlap each other over wider areas. These segments tend to triangulate with each other, forming TGs on both sides. I have some of these TGs which include over 50 shared segments. Since the shared segments triangulate with each other, this is a good pile-up. These TGs are large because more people have these shared segments – probably because the Common Ancestors had large families in Colonial America, leaving us with many, many cousins. Another reason could be a more distant Common Ancestor, who would also leave us a large number of cousins.
In some cases we can use this observation to our advantage. I have a 2nd cousin, on his mother’s side, who is also an 8th cousin, on his father’s side. Our close Common Ancestor was an immigrant to the US in the mid-1800s, and I get relatively few Matches on the segments I share with him. However, on one segment, we have many Matches – it turns out our Common Ancestor is on his father’s side. The tip-off should have been the size of the TG (measured by the number of Matches).
Another observation about large pile-ups…. They will get larger. The number of folks taking an atDNA test is about doubling every 12 months. A consequence of this is that all of our TGs will also double in the next 12 months. So, if you have pile-ups now, they will about double by this time next year. Use these larger TGs to your advantage – work with the Matches to investigate place/time matches, if a Common Ancestor is not easily determined.
Summary
- In general, don’t work with shared segments below 5cM. Most are IBS/IBC – even if they appear to triangulate. We don’t have a good test below 5cM to indicate IBD.
- Watch for, and avoid, pile-ups in the 5-10cM range. These are characterized by many shared segments in the 5-10cM range in a very tight location- usually only 10 or 11cM wide. Move on to larger shared segments in other locations.
- Embrace the large pile-ups. They may from Common Ancestors with large families and/or more distant Common Ancestors. In either case, work with the Matches in these TGs as a Team to determine the Common Ancestor.
18 Segment-ology: Pile-ups by Jim Bartlett 20151007