Endogamy Part II – One Segment from One Ancestor
Review
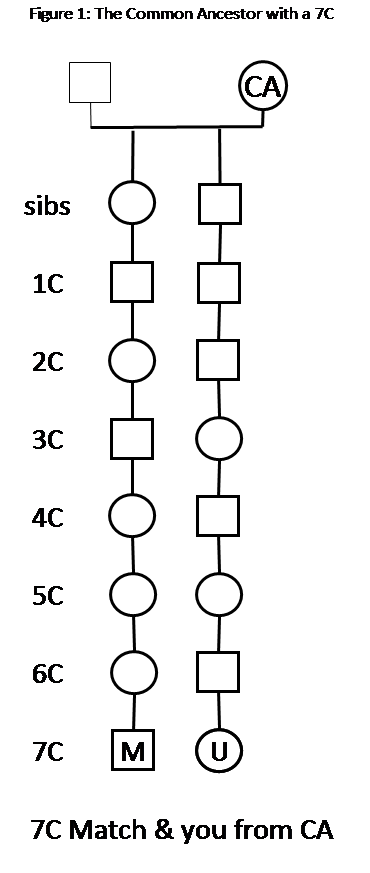
In Endogamy Part I (Shared DNA), we found that the total cM shared between you and a Match is multiplied by the number of times you had the Common Ancestor (CA) in your Tree. So if you and your Match were 5th cousins (5C), you would normally share 3.4cM. If your CA (between you and your Match) was in your Tree five times you would tend to share, on average, 5 x 3.4cM = 17cM. If your Match has that CA in her Tree three times, the total you would tend to share, on average, would be 3 x 5 x 3.4cM or 51cM. For close-cousin Matches (say 1C, 2C, 3C), this make a big difference. For distant-cousin Matches (say 6C-8C or more), where you would probably not match at all most of the time, the endogamy may increase the total cM enough that you’ll actually get an above-threshold Match. Note: a 7C would normally share 0.1cM, so even with an Endogamy factor of E15, you’d only share 1.5cM, on average, and you’d need to be on the tail of the distribution curve to exceed a 7cM threshold for a Match.
In this post – Endogamy PART II – we’ll look at what happens to an individual segment.
Ground Rules
Shared Segment means an IBD segment, from an Ancestor. I usually consider all shared segments over 7cM in a Triangulated Group to be IBD segments.
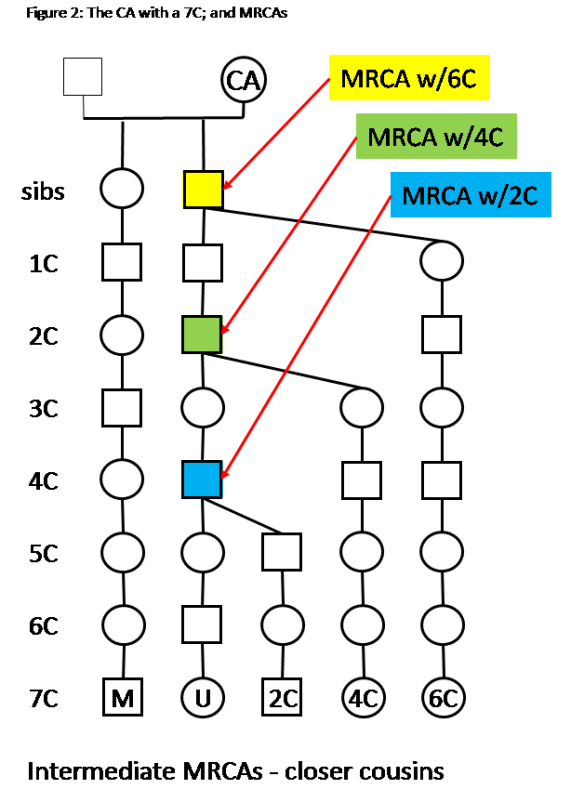
One Ancestor or one CA means one Ancestral line. See CA and MRCA for a discussion of the CA for a shared segment. Note: at different cousinship levels, there may be intermediate CAs, or MRCAs, which are all in one Ancestral line. The shared segment comes from the most distant CA of that Ancestral line. In other words, all the Matches with shared segments (think a Triangulated Group), will have a CA with you on one Ancestral line – you and they will all descend from one CA.
Endogamy means a CA is in our Tree multiple times. So Ancestor A can be represented by A1, A2, A3, etc. for each time that Ancestor is in out Tree. As you read on, you’ll note that it’s important to treat each one (A1, A2, A3, etc.) as a separate Ancestor (even though they are all the same individual).
Assumption: Each Match will have one CA with you. I know in many cases a Match may share multiple Ancestors with you. But for the purposes of this blog post, we will only look at the effects of one CA. We have to build up, one concept at a time. Learn the concept in this post.
Shared Segments from Duplicate Ancestors – analysis
Let’s look at a shared segment between you and a Match. It could be any shared segment. Just to add some reality to this discussion, let’s say it’s on Chr 10 from 8 to 20Mbp with 23cM. We’ll call this SEG-1.
In this example, the CA for you and a Match is in your Tree twice: A1 and A2. Both A1 and A2 are the same person, so both A1 and A2 have the same SEG-1. Let’s look at Figure 1 and see how far SEG-1 can descend toward you.
 Analysis of Figure 1:
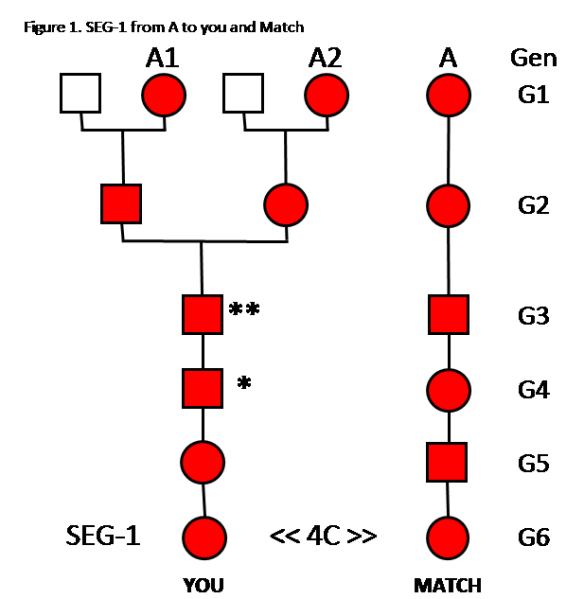
Analysis of Figure 1:
In Generation 6 (G6), YOU and a MATCH are 4C, and share SEG-1. SEG-1 came from Common Ancestor A. Red indicates that person has SEG-1.
In G1, your 2 ancestors, A1 and A2, have SEG-1; and your Match’s ancestor, A, has SEG-1. These are all the same individual – so naturally she has SEG-1, no matter where she is in your Tree or your Match’s Tree.
SEG-1 is passed down from A to the Match through one ancestral line.
In G2, SEG-1 is passed from A1 to her son; and from A2 to her daughter.
In G3 A1’s son passes SEG-1 to the paternal chromosome in his son; and A2’s daughter passes SEG-1 to the maternal chromosome in her son. This G3 son now has two copies of SEG-1, one on his paternal Chr 10, and one on his maternal Chr 10. This is indicated by **.
In G3, the ** father, recombines his two Chr 10s to make one Chr 10 to pass to his son. Only one of the two SEG-1s can be passed on. The G4 son will only get one SEG-1. This is indicated by *. We don’t know whether the maternal or paternal SEG-1 is passed on – it’s a 50/50 chance for either. If you need a refresher on recombination and how only one area of two chromosomes can be passed down, please review Segments: Top-Down.
In G4 and G5, SEG-1 will be passed down to you, and you will share this segment with your 4C Match.
Starting with G1, there are lots of possibilities, but for you and your Match to both share SEG-1, it has to start in a CA (in this case A, A1 and A2) and be passed down through each generation to you and your Match.
One Segment from One Ancestor
This is a fundamental concept of genetic genealogy. Each shared segment can come from only one Ancestor. This means from only one of several Ancestors (A1, A2, A3, etc.) in Trees with endogamy. This can be extended to each Triangulated Group can come from only one Ancestor. A corollary is that a different shared segment (or TG) can come from a different Ancestor. With an Ancestor in your Tree 5 times, it is possible for you to have a different segment from each one. It’s also possible for one Ancestor to pass down several different shared segments (in different TGs). So although we can say “One Segment is from One Ancestor”, the reverse is not true. We have to say “One Ancestor can pass down Multiple Segments” (or no segment).
Shared Segments from Multiple Ancestors – analysis
We can apply this same concept – One Segment from One Ancestor – to even greater endogamy. See Figure 2.
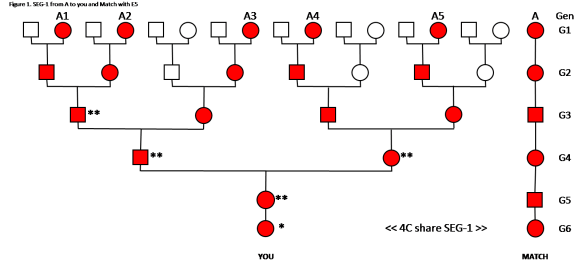
Analysis of Figure 2:
In this case we have E5, the Common Ancestor is in your Tree five times. When cousins (descending from the CA) marry, their child may get two SEG-1s – one on each chromosome. This is indicated by the double **. The next generation gets only one SEG-1 from that line. As noted with the A3 line, the son in G4, could have double ** – one from A1 or A2 (paternal chromosome) and one from A3 (maternal chromosome). The daughter is G5 got a paternal SEG-1 from A1, A2 or A3, and a maternal SEG-1 from A4 or A5. You got a SEG-1 from A1 or A2 or A3 or A4 or A5. At this point we cannot tell which of your Ancestors passed down the SEG-1, but we do know it could only have come from one of them.
In this discussion, I’ve used the “worst case” scenario – each of your A Ancestors passed SEG-1 down as far as she could. In fact, SEG-1 is subject to the 50/50 rule – half the time it will be passed down, half the time it won’t. However, the fact that we share SEG-1 with a Match, and have a Common Ancestor A with that Match, means at least one of your Ancestors A had to pass it all the way down to you and the Match.
We are all well aware that you and a Match probably have multiple Common Ancestors. Again, this analysis does not sort out which Ancestor is the correct Common Ancestor for SEG-1, nor does it sort out which one of the multiple Common Ancestors it is. This analysis just establishes the point that:
One Segment comes from One Ancestor
A very unusual exception
In the case where your parents are cousins, it is possible (but not very probable), that you would carry two SEG-1s (from this example) – one paternal, one maternal. This would be the case in Figure 2 if you were the daughter ** in G5. At GEDmatch, any such segment areas would be highlighted with their “Are Your Parents Related?” tool. So it’s easy to check for such segments, and be aware of where they exist (Chromosome and Start/End locations). In all other cases you don’t need to worry about this issue. For any shared segments meeting this very unusual criteria (exactly the same segment on both chromosomes), you wouldn’t know which of your two ancestors it came from. If there were any difference in these two shared segments (they were not exactly the same), then chromosome mapping would usually tell you which one was which.
Summay:
Each shared segment comes from one Ancestor. In the case of endogamy with multiple identical Ancestors, each shared segment comes from only one of them.
It’s possible to have ten identical ancestors in your Tree, and to get a different shared segment (as in a different TG) from each of them.
16E Segment-ology: Endogamy PART II – One Segment from One Ancestor by Jim Bartlett 20160104