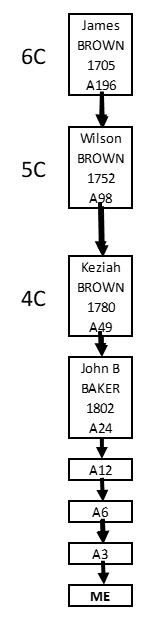
The segment Triangulation Mantra is: 3 separated cousins match each other on the same DNA segment – don’t use close relatives or segments below 7cM.
Besides the fact that this Mantra works, let’s back up a little bit and review why it works – what are we really trying to do. Well… we are really trying to insure all the segments in a Triangulation are from the same side – from just one of your parents. We have two of each chromosome – one from our mother and one from our father. We want our segment Triangulated Groups (TGs) to be on one just one of these chromosomes – to insure we are looking for a Common Ancestor on one side of our Tree (and to insure we don’t intermix the data from our two chromosomes to create a false segment in our TG).
What we want are shared DNA segments on only one of our chromosomes! On one side!
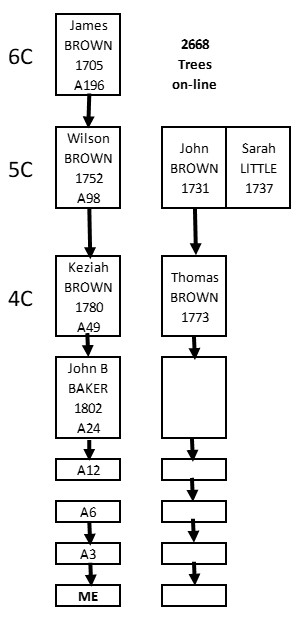
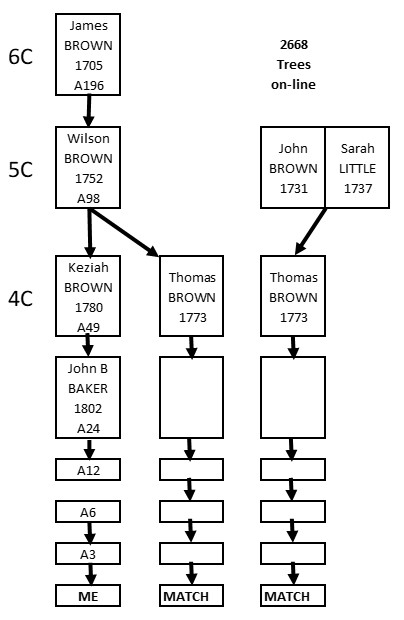
So…. What if we know some of our Matches are on one side – each one is definitely on our maternal or paternal side? Our segment Triangulation becomes almost trivial. Overlapping segments on the same side are Triangulated.
If we have DNA tested a parent, we can relatively easily determine which Matches are on that side. For these, all that is required is segment overlap for Triangulation.
If our parents have significantly different ethnicities, we can often separate many Matches into maternal and paternal sides. Segment overlap with either side equals Triangulation.
Shared Matching (ICW) groups which are clearly one side or the other allow Triangulation just by overlapping segments. This is my preferred method for Triangulation at FamilyTreeDNA (overlapping segments and clear ICW grouping/Matrix). A subset of this is to use the LEEDs method to determine the four grandparents (and thus maternal and paternal groups) and extend to many other Matches who clearly have a Shared Match (ICW) consensus on one side or the other. Use “dots” or Notes to indicate maternal or paternal (or both) sides – these indicators absolutely help identify the side for more Matches.
Clustering often results in Clusters which can be clearly identified as maternal or paternal – this usually applies to all the Matches in each Cluster (but use caution with endogamy).
Note in all of the above, absolutely include parents, aunts/uncles in the TGs! Children and grandchildren should not be included, unless you’ve done other, detailed, analysis on their segments – in general, your descendants don’t add any value except in rare cases when comparing across company lines.
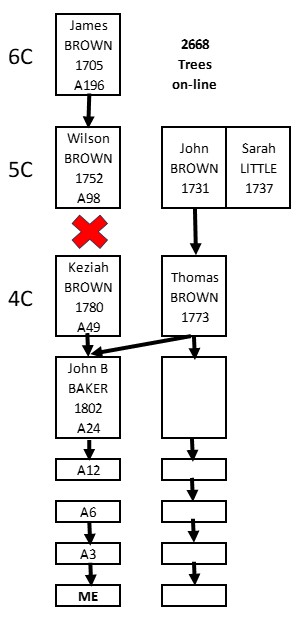
If in doubt – leave it out! Our ancestry (and DNA) sometimes includes some twists and turns. If you come upon something strange, revert to original mantra and check the Match segments against each other.
Take away: If you are sure of the same side, overlapping DNA segments Triangulate.
[11E] Segment-ology: Another Take on Segment Triangulation by Jim Bartlett 20230910